Cart (0 Items)
Your cart is currently empty.
View Products
Reduce the costs of your overlapping peptide library screening experiments without sacrificing data quality. With ProteoGenix’s cost-effective method of peptide library synthesis, you can now employ high-quality peptides for a fraction of the cost. Learn how overlapping peptides can support your research, and reach out to our experts to receive a personalized quote.
If we do not succeed in synthesizing a peptide from your library, you don’t pay for it!
All the peptides synthesized are controlled by MS and HPLC.
We adapt the design and the purity depending on your project to provide you the best data at the best price!
Get 500 peptides in less than 1 month!
With 200000+ peptides synthesized and 99+ % success rate, trust our unrivalled expertise.
We offer an unlimited range of modifications.
Peptide libraries are collections of synthetic peptides, invaluable for studying protein-protein interactions. Through screening and profiling applications, libraries help researchers determine relevant antigen-antibody recognition sites, unravel enzyme-substrate specificity, or determine the sites responsible for receptor binding – particularly useful when studying pathogenic viruses.
Overlapping peptide libraries consist of numerous peptide fragments with the same length and overlapping sequences, spanning a specific segment of a known protein.
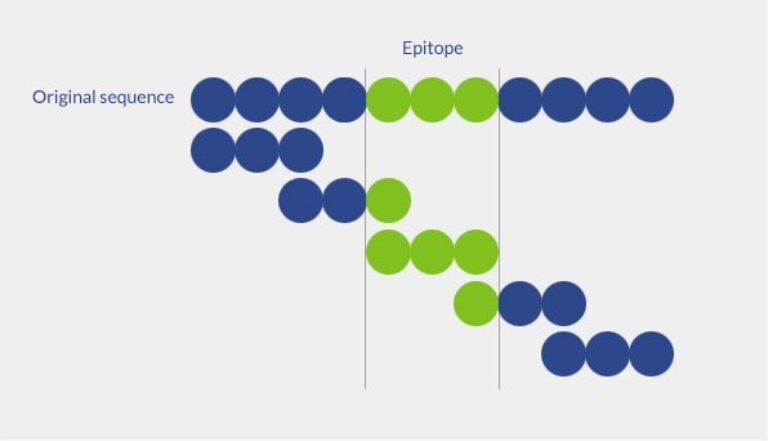
The optimal length of overlapping peptides depends on the intended application. For instance, in epitope mapping experiments, peptides should contain between 9 and 22 amino acids because epitopes typically fall within that size range.
In contrast, offset numbers are less application-dependent, reflecting the degree of overlap within the library. In practice, this parameter is defined by the number of amino acid residues shared by neighboring peptides. Ideally, this number should be 1/3 of the full length of the peptide fragments, to ensure all regions of the protein segment are equally covered.
By defining these two parameters, overlapping peptide libraries can be easily designed using ProteoGenix’s free Peptide Library Generator tool:
The ratio between length and offset dictates the size of the overlapping peptide library and thus determines the quality of the data derived from the experiment. Although smaller libraries are much easier and less expensive to synthesize, the likelihood of researchers finding the precise interface residues in complex projects is often limited. As a result, keeping costs low may be detrimental to the quality of the data generated by these assays.
When offset numbers are high and peptides long, the resulting library will have fewer peptides. In this case, although researchers will likely encounter a hit, it will be exponentially harder to pinpoint the specific interface residues.
In contrast, when offset numbers are low and peptides short, the resulting library will have more peptides, making it easier to identify the interface residues but also resulting in more time-consuming experiments.
To overcome this limitation, researchers often group different peptides into pools. In this case, instead of screening hundreds of peptides individually, researchers can focus on identifying positive peptide pools.
Two major methods have been developed to effectively screen peptide pools:
Want to learn more about how we can support your project? Generate your overlapping peptide library for free and reach out to our experts today:
Overlapping peptide libraries are extremely useful for identifying linear or continuous regions that can interact with a particular monoclonal antibody, enzyme, or receptor. This is because peptides, particularly short peptides, lack tertiary structures and thus cannot replicate native antigen conformations or discontinuous epitopes.
Nevertheless, these libraries serve numerous applications, including:
Epitope mapping experiments are typically performed by ELISA, whereas overlapping peptides are biotinylated and coated on a microtiter plate. Subsequently, enzyme-conjugated antibodies (typically with horseradish peroxidase, HRP) and chromogenic substrates are used to detect positive antibody-epitope interactions.
In theory, all enzymes can be profiled through overlapping peptide libraries. However, only assays to detect kinase and protease activity have been developed. Immobilizing peptide substrates on a membrane is a standard strategy for capturing kinase activity. After immobilization, the medium is supplemented with ATP, and kinase-substrate binding specificity is assessed by autoradiography. On the other hand, protease-substrate recognition is detected by conjugating peptides with fluorescent dyes. Proteases cleave the tag after specified reaction times, allowing for accurate activity quantification.
In contrast, T-cell studies require whole cells. Unraveling key T-cell epitopes is vital for vaccine and immunotherapy development. During the immune response, T-cells only recognize antigenic fragments non-covalently bound to major histocompatibility complex (MHC) proteins on the surface of antigen-presenting cells (i.e., dendritic cells, macrophages, B-cells).
Multiple studies demonstrated that synthetic peptides conjugated with molecules such as insulin, phytohemagglutinin (PHA), or purified protein derivate (PPD) from Mycobacterium tuberculosis can also trigger the same response in vitro. T-cell interactions can then be analyzed using conventional ELISPOT assays.
Start your project today, by generating your overlapping peptide library for free.
You could also be interested in:
