Cart (0 Items)
Your cart is currently empty.
View Products
Need to analyze hundreds of peptides? Save time and let us synthesize your peptide library. Thanks to our state-of-the-art peptide synthesis platform, we can deliver a huge number of high quality peptides (including modifications) directly on plates for various applications: proteomics, epitope mapping, cancer therapy, therapeutic antibody development…
If we do not succeed in synthesizing a peptide from your library, you don’t pay for it!
All the peptides synthesized are controlled by MS and HPLC.
We adapt the design and the purity depending on your project to provide you the best data at the best price!
Get 500 peptides in less than 1 month!
With 200 000+ peptides synthesized and 99+ % success rate, trust our unrivalled expertise.
We offer an unlimited range of modifications.
Peptide libraries consist in a large collection of peptide sequences used to determine to critical fragments for function or binding. Peptide libraries are used in a wide variety of applications such as proteomics, vaccine development, epitope mapping or cancer therapy. Compared to peptide phage display, peptide libraries offer the possibility to screen modified peptide and peptides containing unnatural or D-form amino acids.
Overlapping peptide libraries are commonly used for linear epitope mapping. The goal of this technique is to generate a library of overlapping peptide sequences. Each peptide libraries is defined by two parameters: the peptide length and the peptide offset meaning that each peptide of the library will be characterized by a same peptide length but by a different primary structure depending on the offset. The peptide offset defines the number of overlapping amino acid residues between 2 adjacent sequences and corresponds to the degree of overlap.
The two parameters are generally chosen by considering two factors: the cost and the value of the data (and the risk of missing the epitope). For example, for linear epitope mapping:
Overlapping peptide libraries can be used and adapted for several applications such as protein-substrate interaction screening, T-cell epitope discovery, T-cell stimulation…
Parameters to consider: peptide sequence, peptide length, peptide offset.
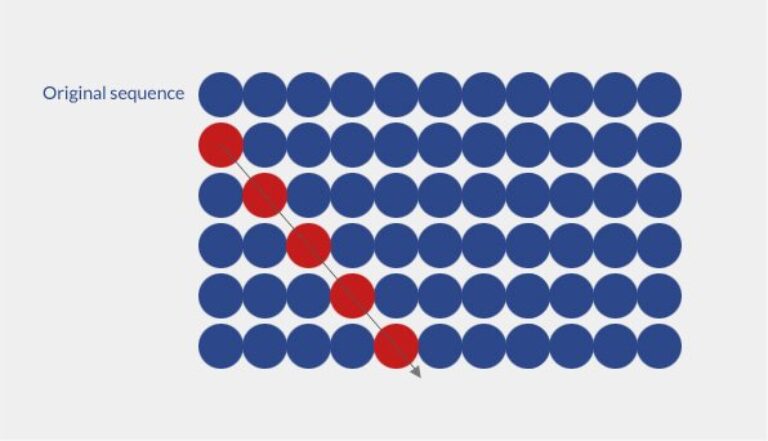
Alanine scanning libraries are specifically designed to determine the contribution of a specific amino acid in the function, activity or stability of a peptide or protein structure. Substitution of a key residue by an alanine results in a reduction of the peptide activity and demonstrates the importance of this substituted residue in the global structure. From a technical point of view, the library is designed by sequential alanine substitution of each non-alanine residues. Alanine is chosen because it is chemically inert and it is the smallest existing chiral amino-acid residue. However, other amino-acids can be used upon request.
Parameters to consider: peptide sequence.
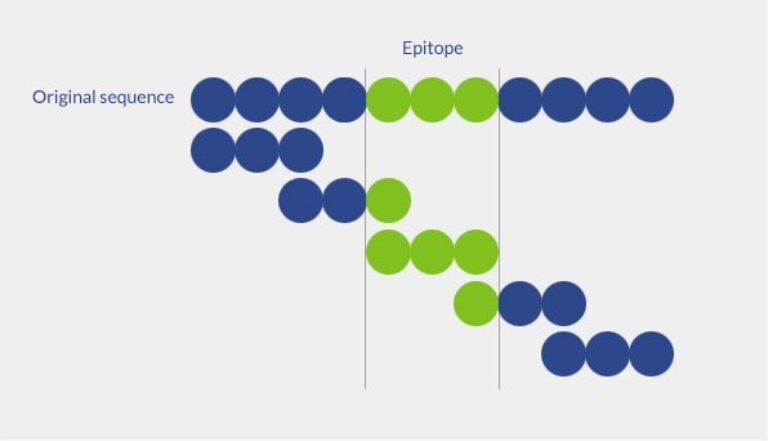
Truncation peptide libraries allow determining the shortest peptide sequence necessary to observe an epitope activity. The peptide library is constructed by systematic removal of terminal amino acids. The truncation step can be adapted following your requirements. Also, if the essential amino acids are already known (eg. by a previous alanine scanning), it is possible to fix the direction of truncation.
Parameters to consider: truncation increment, truncation direction (N-ter, Cter or both if essential amino-acids are known).
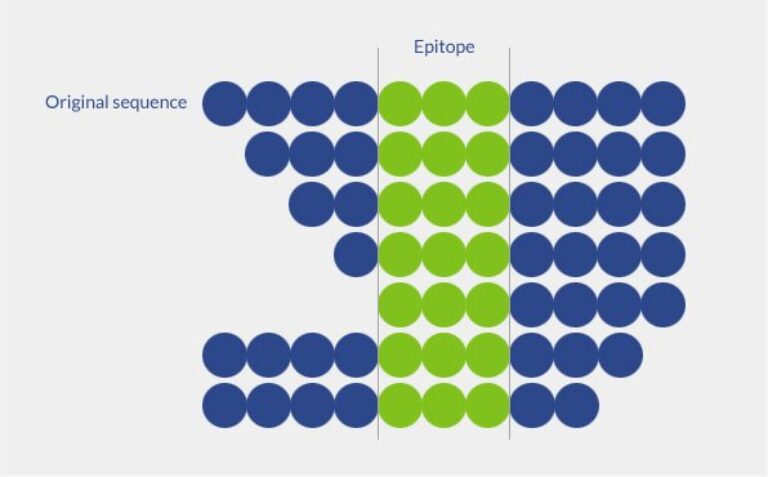
Positional scanning libraries are particularly useful for peptide sequence optimization. Amino acid residues at defined positions are replaced by all other amino-acids, one at a time. Then, activity measurement allows determining which amino-acid residue increases the peptide activity. Positional scanning libraries can also be useful to improve T-cells or antibody epitopes.
Parameters to consider: peptide sequence, positions to vary.
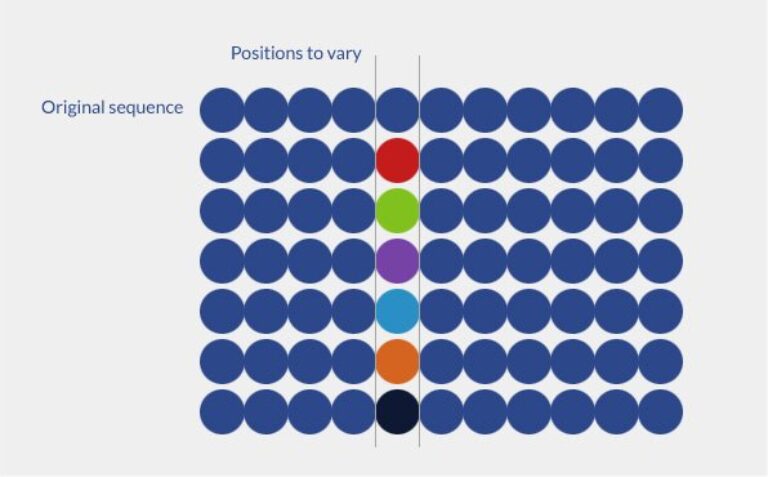
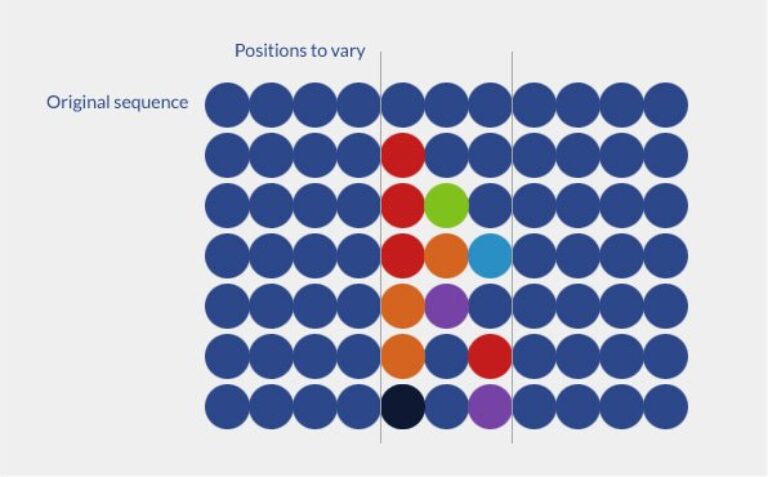
A random peptide library is consists in randomly generating variations of a peptide sequence by varying randomly the amino acid residues within a defined portion of the whole peptide sequence. Random libraries are used to optimize peptide sequence or to generate novel and active peptide sequences.
Parameters to consider: peptide sequence, positions to vary.
A scrambled peptide library is composed of all the alternative sequences of a given peptide. This means that all the amino-acid residues of the initial sequence are mixed to form all the possible amino acids combination. This type of library is usually used to optimize a peptide sequence.
Parameters to consider: peptide sequence.

